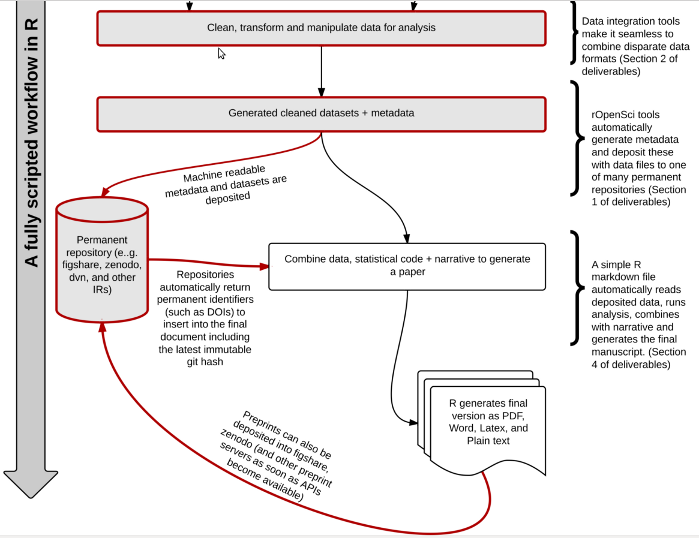
There is confusion between the definitions of reproducibility, repeatability and replicability. I strongly feel we need to tackle that head on and come to an agreed definition. I prefer Peng 2011:
- Reproduciblity is using the same data and getting the exact same result.
- Replication is getting a new sample and doing the analysis again and getting a similar result.
Peng, R. D. (2011). Reproducible research in computational
science. Science, 334(6060), 1226–1227. doi:10.1126/science.1213847
There are many people using these interchangeably or around the opposite way. For example Drummond got it round the wrong way in ‘Drummond, C., 2009. Replicability is not reproducibility: nor is it good science’ http://cogprints.org/7691/7/icmlws09.pdf and then reverted it in ‘Reproducible Research: a Dissenting Opinion’. http://cogprints.org/8675/1/ReproducibleResearch.pdf (Check out Peng’s reaction: http://simplystatistics.org/2012/11/15/reproducible-research-with-us-or-against-us-3/)
And this blog but also gets the definitions around the wrong way http://jermdemo.blogspot.com.au/2012/12/the-reproducible-research-guilt-trip.html (even tho being quite entertaining to read and had this great picture…. not sure what the picture means???)

Sure, OK, it is fine that people define things differently to one another but:
The single biggest problem in communication
is the illusion that it has taken place.
George Bernard Shaw quotes from BrainyQuote.com
We rely on a common defintion to ensure we are talking about the same thing.
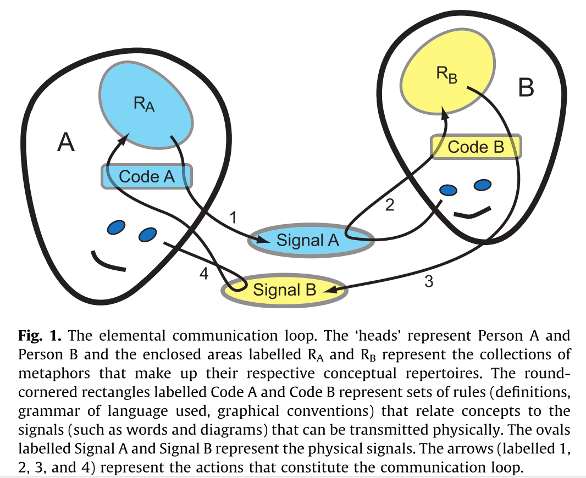
Source: Newell, B. (2012). Simple models, powerful ideas: Towards effective integrative practice. Global Environmental Change, 22(3), 776–783. http://dx.doi.org/10.1016/j.gloenvcha.2012.03.006
It is regrettable that in Ecology (my favourite discipline) there seems to be quite a wide gap between various author’s definitions. In CASSEY, P., & BLACKBURN, T. M. (2006). Reproducibility and Repeatability in Ecology. BioScience, 56(12), 958. http://bioscience.oxfordjournals.org/content/56/12/958.full they agree with the definition of Peng 2011. However another author gives a very confused and overlapping view:
because that context changes through time and space, it is virtually
impossible to reproduce precisely or quantitatively any single
experimental or observational field study in ecology. Yet many
ecological studies can be repeated. In particular, ecological
synthesis – the assembly of derived datasets and their subsequent
analysis, re-analysis, and meta-analysis – should be easy to repeat
and reproduce
Ellison, A. (2010). Repeatability and Transparency in Ecological Research. Ecology. https://dash.harvard.edu/bitstream/handle/1/3123279/Ellison_Repeatability.pdf?sequence=2 Accessed 12 Jan 16
In another interesting approach Freedman, L. P., Cockburn, I. M., & Simcoe, T. S. (2015). The Economics of Reproducibility in Preclinical Research. PLOS Biology, 13(6), e1002165. http://dx.doi.org/10.1371/journal.pbio.1002165 chose instead to define irreproducibility such that it:
that encompasses the existence and propagation of one or more errors,
flaws, inadequacies, or omissions (collectively referred to as errors)
that prevent replication of results
Leaving us to assume that the opposite of this is therefore reproducibility, although avoiding defining this themselves. Looking back at the two heads in the picture above… it is interesting to ponder how some people would receive the signal of Freedman et al, having defined the opposite of the thing that is the object of their discussion, rather than the thing itself!
Let’s all agree with Peng and Cassey/Blackburn and move on already!